selected atom:
none
Activity 2
Exploring Molecular Structure Using MolVisWeb
Adapted from Parthena E. Kotsalidis et. al. (1)
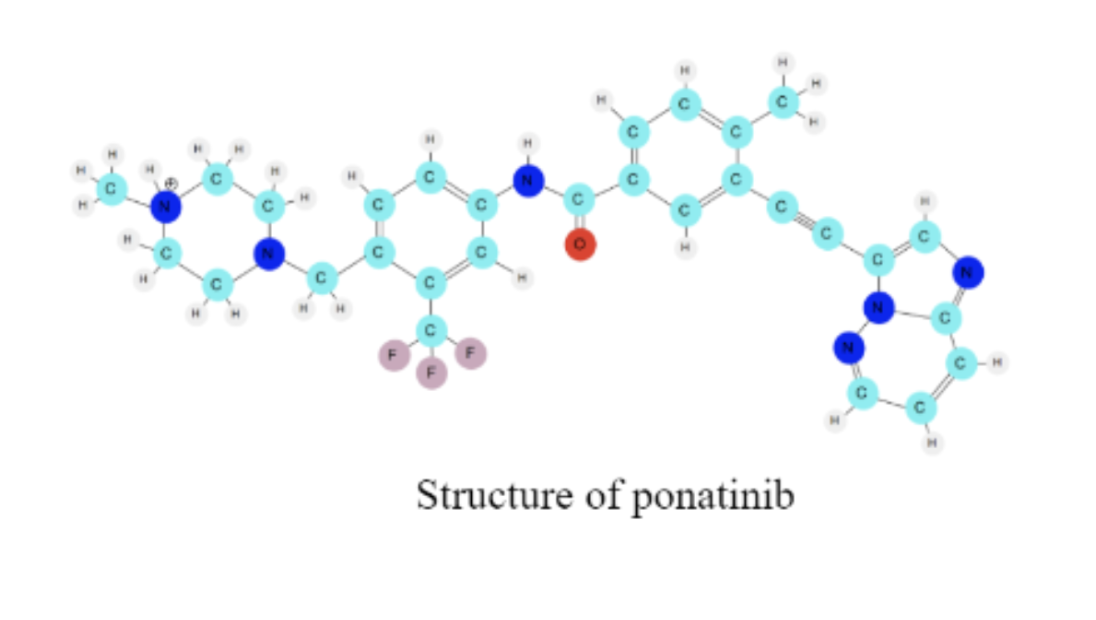
Exploring the Drug Ponatinib
Note: If you would like to access these questions in Google Document format, click on the following link: Activity 2 Google Document.Chronic Myeloid Leukemia (CML) is a cancer of the myeloid cells, which are cells that make red blood cells, platelets and most types of white blood cells.(2) Patients with cancer produce more white blood cells than they need due to the overactive BCR-ABL gene. However, highly effective drugs have been developed to inhibit the function of this overactive kinase in patients with CML. One such drug is Imatinib, which binds to the Abl kinase and inhibits its function. Imatinib works by fitting into the binding pocket of the Abl kinase, the same binding pocket ATP would bind in. When imatinib binds, the Abl kinase can no longer bind the ATP molecule it needs to function. As a result, the overactive kinase gets turned off.
There are many drugs that can be used to treat CML and that can bind to this same protein. The specific way that drug molecules bind to the protein depends on the intermolecular forces (noncovalent interactions) between the molecules. Today, you are going to explore the structure of another drug used to treat CML, called ponatinib. The structure of the drug molecule determines how it will interact with the protein target.
I. Loading the Drug Molecule into MolVisWeb
1. Open the molecular visualization software (you're already here!)
2. Three panels are currently visible: the control panel (left), the molecule display panel (middle), and the activities panel (right). Note that the default molecule displayed is Caffeine.
3. If you would like to, you can hide the activities panel on the right using the button in the upper right corner of the screen.
4. Locate the control panel on the left side of the screen. For the drop-down menu titled “molecule”, switch Caffeine to Ponatinib.
5. The drug molecule should now be loaded.
6. Click your mouse anywhere on the molecular visualization window and hold. Drag your mouse to move the molecule around the screen. In a few words, describe what you see on your screen.
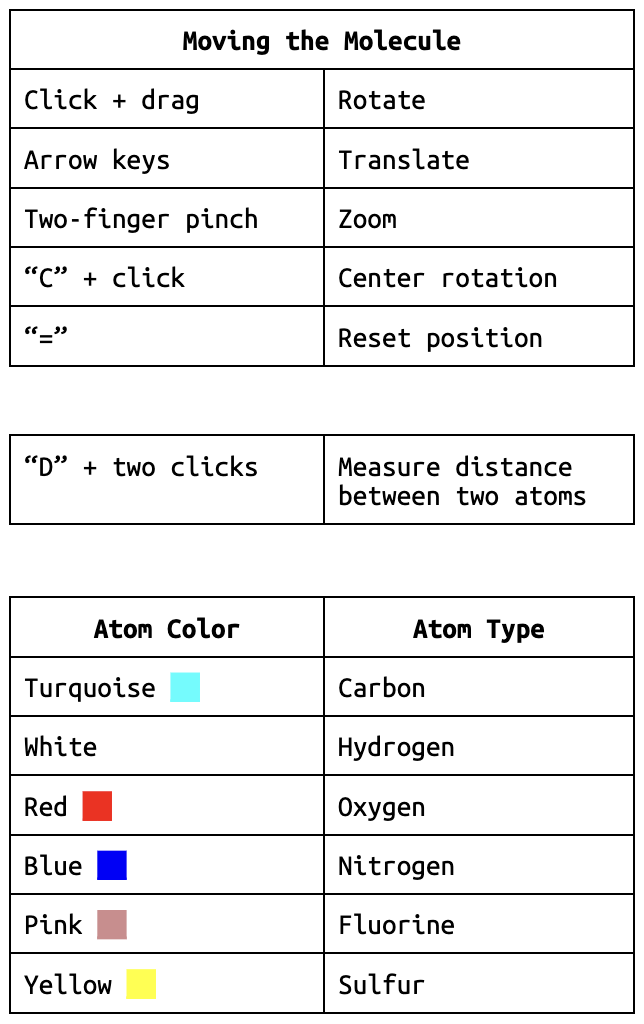
Each atom in the molecule has its own unique color. You can find the atom colors in the help menu indicated by the “?” in the bottom left corner.
7. What one type of atom makes up most of the molecule?
8. Do you think the atoms in the molecule are connected with covalent bonds, ionic bonds or intermolecular forces? Explain your answer.
9. Identify one more structural feature of the molecule that interests you.
II. Viewing the Molecule
★ Tip: You can center the display around the molecule by pressing the “=” key. ★
1. First, let’s explore translating and rotating the molecule. As you discovered before, dragging the mouse around the screen rotates the molecule. What movement occurs when you use the up, left, right, and down arrow keys to move the molecule?
2. Now we are going to explore centering and rotating around a specific atom. Press the “C” key to turn on centering mode. Your mouse icon should turn into a pointing finger.
Pick one atom at one of the ends of the drug molecule and click on it with your mouse. Then press “C” again to exit centering mode and rotate the molecule by dragging with your mouse. Describe how your molecule moves.
★ You can find a summary of the most common MolVisWeb shortcuts and commands in the help menu, located in the bottom left corner of your screen. ★
3. Take a moment to study the drug molecule once again. Use your new skills with rotating, translating, scaling, and centering the molecule to answer the following questions.
3a. How many ring-like structures can you find on the molecule?
3b. Where do you notice that the white colored hydrogen atoms are located on the molecule?
3c. How many red oxygen atoms do you see?
3d. Determine the number of pink colored fluorine atoms on the molecule.
4. List two similarities and differences between a 2-D molecular structure of ponatinib and the 3D molecular structure that you can manipulate on your screen.
II. Drawing Methods
1. In the control panel, notice the parameters that you are currently working with:
Drawing Method: Ball-and-stick
Coloring Method: Name
Next, let’s experiment with a few different Drawing Methods.
2. Input the following selections in the Graphical Representations window, and describe what you see.
Drawing Method: Space filling
Coloring Method: Name
3. Next try the following settings. Describe what you see.
Drawing Method: Lines
Coloring Method: Name
4. Now that we’ve seen a variety of different drawing methods, let’s compare them! Match the following statements with the drawing method that you think would be most helpful in answering the question. More than one drawing method can be used for a single question.
4a. Which drawing method, in your opinion, is most helpful in identifying specific atoms – Lines or Ball-and-stick – and why?
4b. There are many interesting structural features in the ponatinib molecule. One such feature is a triple bond that looks like a longer extended line. Which drawing method, in your opinion, is most helpful in identifying the linear region in the ponatinib molecule – Space filling or Line – and why?
4c. Which drawing method, in your opinion, best helps you determine the angles around the atoms – Lines or Space filling – and why?
4d. Which drawing method, in your opinion, best helps you determine the amount of space taken up by the atoms in the molecule – Lines or Space filling – and why?
4e. Which drawing method, in your opinion, is most helpful in identifying the rings in the molecule – Lines or Space filling – and why?
4f. Do you think there is one “best” way to represent or draw molecules? Why or why not?
IV. Coloring Methods
MolVisWeb offers four different coloring methods. Name, which you’ve already explored, colors atoms by element identity. The other three coloring methods color the entire representation a single color.
V. Determining Distances Between Atoms
1. Input the following parameters into the Graphical Representations Window:
Drawing Method: Ball-and-stick
Coloring Method: Name
2. Press the “D” key. You will see a white cross appear on your screen in place of your mouse.
3. Click on a pink fluorine atom. A label will appear next to the fluorine atom. (Depending on the orientation of the molecule, you may need to rotate it to see the label. Alternatively, the currently selected atom label will show up in the left panel.)
4. Fill in the blank for the label DRG285:F____. (DRG is an abbreviation for drug. There are three fluorine atoms, so each one has a different number 34-36.)
5. Now click on the red oxygen atom. A label will appear next to the oxygen atom.
6. Fill in the blank for the label DRG285:O____.
7. You will see a white line between the two atoms with a green number. The number that pops up is the distance between the two atoms. MolVisWeb measures distance in Angstroms where 1 Angstrom = 10-10 meters. What is that distance?
★ Tip: You can hide labels and bonds by pressing “D” and clicking on the two atoms again. You can also click on the “clear bonds” button at the bottom right of the control panel. ★
8. Pick two other atoms to find the distance between and do so. Write the names of the atoms and their distance.
CHALLENGE

1. Find one central atom with each of the following molecular geometries. (Remember that you can find the label of an atom by clicking on it and looking at the letters and numbers after the colon (e.g., “C31” or “H40”) that show up on the control panel. Again, the “DRG” before the colon stands for “DRUG” and refers to what the molecule has been named in this case.)
Indicate the labels of the atoms you find for the following geometries: Tetrahedral, Trigonal planar, Trigonal pyramidal, Bent, Linear.
Exploring the Protein: Abl Kinase
Thus far, you have learned that imatinib and ponatinib are two drugs that can treat chronic myeloid leukemia. They both bind to a kinase protein called the Bcr-Abl kinase. In addition, you have explored basic ways to use MolVisWeb to manipulate and analyze the structure of the drug molecule ponatinib. Next, you will explore the structure of the Abl kinase portion of this protein. Proteins are large complex molecules that are made up of many smaller units called amino acids. An amino acid in a protein is often called a “residue”. Proteins play a critical role in your body. They are required for regulation of the body’s tissues and organs, and they are required for structure and function.
Loading the Protein: Abl Kinase into MolVisWeb
1. In the control panel, select the Abl kinase molecule from the top drop-down menu. It might take a few seconds to load.
2. Now, the Abl kinase protein is shown in your display panel.
Use the skills you have learned thus far to explore the molecule.
3. List some of the different types of atoms you see in the protein.
II. Drawing Methods
As you observed in the earlier exploration with the drug, many valuable methods exist for representing molecules.
Notice the parameters that you are currently working with:
Drawing Method: Ball-and-stick
Coloring Method: Name
1. There are three Drawing Methods in MolVisWeb: Space filling, Ball-and-stick, and Lines. Spend a few minutes exploring each method with the protein.
2. Experiment with a few of the different Drawing Methods and decide which methods you like the best for the protein. List your favorite Drawing Methods.
3. Now that you’ve explored a variety of different drawing methods let’s compare them! Match the following statements with the drawing method that you think would be most helpful in answering the question.
3a. Which drawing method, in your opinion, better helps you determine the amount of space taken up by the protein – lines or space filling? Explain your answer.
3b. Do you think there is one “best” way to represent or draw protein molecules? Explain your answer.
III. Amino Acid Residue Selections
As mentioned previously, an amino acid (small units that make up proteins) within a protein can also be called a “residue”. There are 20 amino acid residues used to make proteins. The order these amino acids string together determines the primary sequence of the protein, which in turn can determine how the protein folds into a 3D structure.
The Abl kinase protein you are working with today has a unique 3D structure determined by the sequence of amino acid residues that make it up. It has a total of 284 amino acids. The Amino Acids Reference Sheet shows the structural formulas for the amino acids.In the control panel enter the following:
Drawing Method: Ball-and-stick
Coloring Method: Name
In the “SELECTION METHOD” section, make sure you’re in the “Residue” tab. Replace the word “all” with “86”. Press enter to apply your changes.
You’ve just selected an amino acid residue.
★ Tip: You can recenter the view of the molecule by pressing the “=” key.★
1. Use your Amino Acids Reference sheet to write the name, three letter code, and single code for the amino acid you selected.
2. Rotate, translate, and scale the 3D amino acid on your screen. In what ways does the 2D structure of the amino acid on your reference sheet compare to the 3D structure of amino acid on your MolVisWeb screen? List two similar and two different properties.
3. Now in the “Residue” tab text box type “210”. Press enter on your keyboard again and center and zoom as needed using the commands you have learned. As a reminder, you can also press “=” to center on your newly selected residue.
You’ve just selected another amino acid residue.
4. Use your Amino Acids Reference sheet to write the name, three letter code, and single code for the new amino acid, resid 210, you selected.
5. Rotate, translate, and scale the 3D resid 210 on your screen. In what ways does the 2D structure of the amino acid on your reference sheet compare to the 3D structure of amino acid on your MolVisWeb screen? List two similar and two different properties.
IV. Protein Structure
Proteins like the Abl kinase contain amino acids. Amino acids have a unique structure. Each amino acid shares a set of atoms that make up the amino acid backbone. The central carbon atom, also called the alpha carbon, has an atom or a group of atoms attached to it that varies among the amino acids (These groups are called side chains). When strung together, these amino acids make up a protein.
Here’s what a string of amino acids looks like:
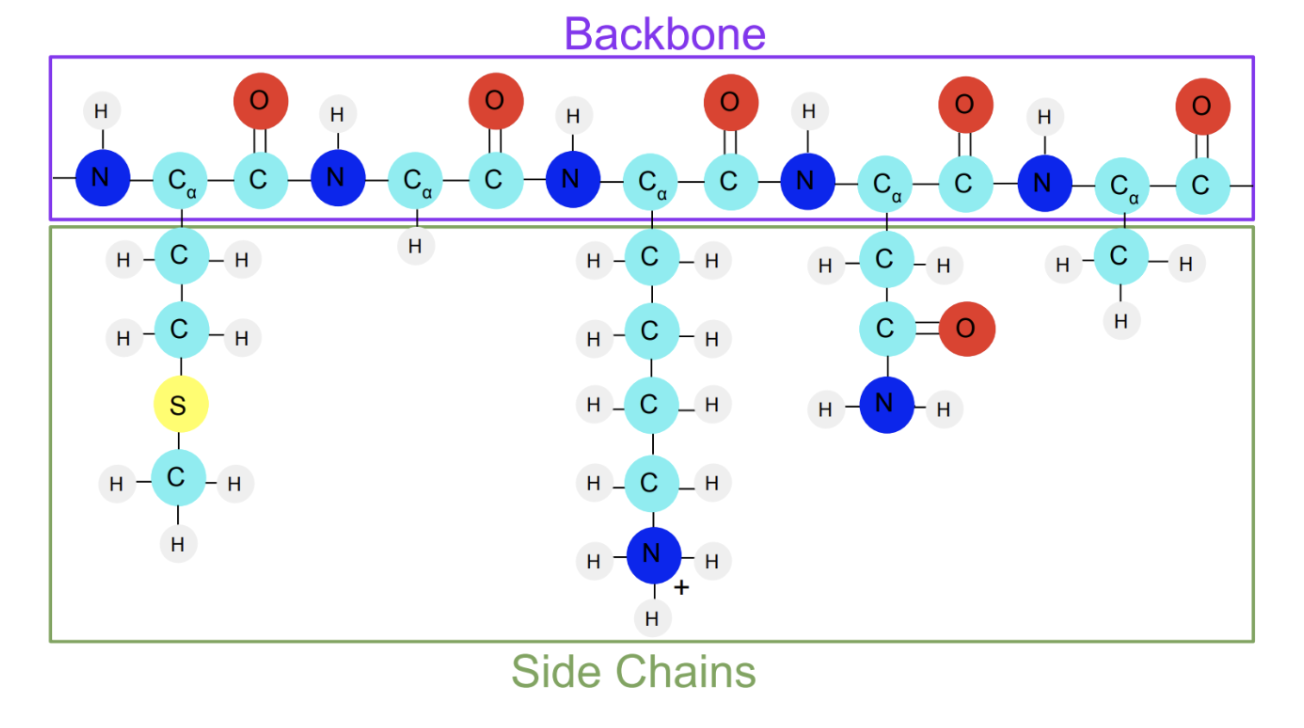
Next, let’s look at the backbone of the Abl kinase protein.
1. In the “SELECTION METHOD” section, select the tab “Molecule”.
2. In the “molecule” box, type “backbone”. Press enter.
3. Describe what you notice about the backbone of the Abl kinase protein.
4. What types of atoms do you see in the backbone? List the types of atoms in the space below.
5. Based on what you see in the representation, why do you think it is called the “backbone” of Abl Kinase?
END
Congratulations! You have just finished manipulating, analyzing, and exploring your first molecules using MolVisWeb. Click "next" to continue on to the next activity.
Sources
(1) Kotsalidis, P. E.; Kranc, S. N.; Berryman, M.; Radhakrishnan, M. L.; Elmore, D. E. EMMAs: Implementation and Assessment of a Suite of Cross-Disciplinary, Case-Based High School Activities to Explore Three-Dimensional Molecular Structure, Noncovalent Interactions, and Molecular Dynamics. J. Chem. Educ. 2024, 101 (6), 2436–2447. https://doi.org/10.1021/acs.jchemed.4c00036.
(2) “What Is Chronic Myeloid Leukemia?: Leukemia Types.” American Cancer Society, 19 June 2018, https://www.cancer.org/cancer/chronic-myeloid-leukemia/about/what-is-cml.html.